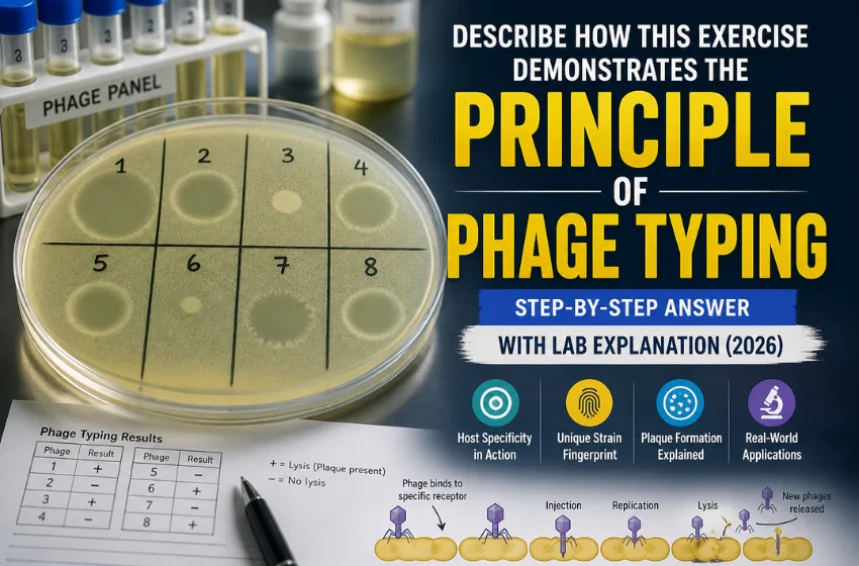
You’ve seen the petri dishes. You’ve watched the clear patches appear on a bacterial lawn. And now you need to put it all into words: describe how this exercise demonstrates the principle of phage typing.
Here’s the thing: most explanations get either too technical, too fast, or too shallow to actually be useful for coursework, exams, or real lab understanding. This guide hits the middle ground. Whether you’re a microbiology student working through an assignment, a lab professional refreshing foundational knowledge, or a curious reader who stumbled onto one of the most elegant techniques in all of infectious disease science this covers everything you need, clearly and completely.
We’ll walk through what phage typing actually is, why the exercise works the way it does, what the results mean, and how this lab demonstration connects to real-world outbreak investigations, hospital infection control, and the cutting edge of 2026 microbiology research.
What Is Phage Typing? The Core Principle in Plain
Before you can describe how an exercise demonstrates a principle, you need to understand the principle itself.
Phage typing is a method used to identify and classify bacterial strains based on their susceptibility or resistance to a panel of specific bacteriophages. A bacteriophage (or “phage”) is a virus that infects bacteria. Not all bacteria. Specific bacteria.
That specificity is the entire point.
Think of it this way: every bacterial strain has a unique set of surface receptors molecular “locks” on its outer wall. Bacteriophages carry matching “keys” surface proteins called tail fibers that recognize and bind to those specific locks. If the key fits, the phage attaches, injects its genetic material, hijacks the bacterium’s cellular machinery, replicates itself, and eventually causes the bacterium to burst open. That bursting is called lysis.
If the key doesn’t fit, nothing happens. The phage moves on. The bacterium survives.
This lock-and-key specificity between phage and host bacterium is the principle that phage typing exploits. By exposing an unknown bacterium to a standardized panel of known phages and observing which ones cause lysis, scientists can generate a phage type profile essentially a fingerprint that identifies the bacterial strain.
The core principle in one sentence: Because each bacteriophage can only infect bacteria with matching surface receptors, the pattern of lysis or resistance across a panel of phages uniquely identifies a bacterial strain.
A Brief History: Where This Principle Came From
Understanding the history makes the exercise make more sense.
The first use of bacteriophages to identify bacteria was reported in 1925, when Sonnenschein used typhoid and paratyphoid phages to diagnose typhoid fever. A decade later, in 1934, researchers discovered that some strains of Salmonella typhi displayed surface structures called Vi antigens and that certain phages could only lyse strains carrying those antigens. That was the first documented demonstration of what we now call phage typing: using phage-host specificity to distinguish between bacterial strains.
By 1938, Craigie and Yen had refined the approach, using Vi phages at carefully calibrated dilutions to differentiate 11 distinct types of Bacillus typhosus. The method spread rapidly. Felix and Callow extended it to Salmonella paratyphi B in 1943. The International Committee for Enteric Phage Typing was established in 1947, standardizing the method globally.
For decades, phage typing was the gold standard for bacterial strain identification in outbreak investigations from tracking Salmonella through food supply chains to tracing Staphylococcus aureus between hospital wards.
Molecular methods have since added new layers of precision, but phage typing remains foundational and the lab exercise that teaches it is still one of the clearest demonstrations of a core biological principle anywhere in microbiology education.
See Also : Droven io AI Technology, IT Services & Business Automation Platform for the USA
The Exercise, Step by Step: What You Actually Do
Here is the standard phage typing lab exercise, broken down clearly so you can both understand it and describe it accurately.
Step 1: Prepare the Bacterial Lawn
A susceptible bacterial culture often Staphylococcus aureus or Salmonella typhi in teaching settings is spread evenly across the surface of an agar plate. This creates what microbiologists call a bacterial lawn: an unbroken layer of growing bacteria covering the entire plate surface.
The lawn is the canvas. Without it, you have nothing to observe.
Step 2: Apply the Phage Panel
The agar surface is divided into a grid typically using a template placed under the plate. Each grid square is inoculated with a drop of a different, known bacteriophage suspension. In standardized phage typing schemes, these phages come from an internationally recognized collection. In teaching exercises, a smaller panel of phages (often 3–10) is used to demonstrate the principle.
Each phage is applied at its critical test dilution (RTD) a carefully established concentration that produces reliable, reproducible results. Too concentrated, and every phage seems to kill everything. Too dilute, and nothing grows. The RTD is the sweet spot where specificity becomes visible.
Step 3: Incubate
The plate is incubated at the optimal growth temperature for the bacterium typically 37°C for human pathogens for 18 to 24 hours. During this time, the bacterial lawn continues to grow and the phages interact with the bacteria in each grid square.
Step 4: Observe Plaque Formation
After incubation, you examine the plate. In some grid squares, the bacterial lawn appears completely intact, a uniform cloudy surface showing continued bacterial growth. In others, you see plaques: clear zones where bacterial cells have been lysed and destroyed by the phage.
A plaque is visible evidence of successful phage infection. Every clear zone represents millions of individual infection-lysis cycles, each one demonstrating the lock-and-key specificity at the heart of the principle.
Step 5: Record and Interpret the Pattern
The pattern of which grid squares show lysis (plaques present) and which show intact growth (no plaques) is the phage type profile of that bacterial strain.
This pattern is then compared against a reference chart. The combination of positive and negative results which phages lysed the bacteria and which did not map the bacterium to a known phage type.
How the Exercise Directly Demonstrates the Principle
Now we can answer the question directly.
The exercise demonstrates the principle of phage typing in four specific, observable ways:
1. It Makes Host Specificity Visible
The principle states that each bacteriophage infects only bacteria with specific matching surface receptors. The exercise proves this by showing that only certain phages out of the entire panel applied cause lysis. The bacterial lawn survives intact under phages that do not match its surface receptors, and is destroyed only where a matching phage was applied. You can see host specificity happening in real time, in physical space on the agar surface.
2. It Demonstrates That Lysis Is Selective, Not Universal
If phages destroyed all bacteria indiscriminately, every grid square would show a plaque. But that’s not what happens. The pattern of lysed and unlysed squares is specific to the bacterial strain. Identical phages, applied to a different bacterial strain on a parallel plate, produce a different pattern. This confirms that the interaction is determined by the bacterium’s surface characteristics not by random chance or general phage aggression.
3. It Creates a Reproducible, Identifiable Fingerprint
The exercise demonstrates that the phage susceptibility pattern is stable and reproducible for a given bacterial strain. If you repeat the experiment with the same strain, you get the same pattern. If you use a different strain, you get a different pattern. This consistency is what makes phage typing scientifically useful; it transforms the biological principle of specificity into a practical identification tool.
4. It Connects Molecular Mechanism to Observable Outcome
Students cannot directly observe receptor-ligand binding or phage DNA injection. But they can observe the downstream consequence of those molecular events: the plaque. The exercise bridges the gap between the invisible molecular mechanism (phage tail fiber binding to bacterial surface receptors) and the visible macroscopic result (clear zone in the bacterial lawn). This cause-and-effect chain is the principle made tangible.
Understanding Plaque Formation in More Detail
A plaque is not just a “clear spot.” Understanding what it actually represents makes the principle clearer.
When a phage successfully infects a bacterium, it initiates the lytic cycle:
- Adsorption The phage’s tail fibers recognize and bind to specific receptors on the bacterial cell surface. This is the step governed entirely by surface specificity.
- Injection The phage injects its nucleic acid (DNA or RNA) into the bacterial cell.
- Replication The bacterial cell’s machinery is redirected to produce hundreds of new phage particles.
- Lysis The bacterial cell bursts open, releasing the new phages into the surrounding environment.
- Propagation The released phages infect neighboring bacteria, creating an expanding zone of lysis.
The visible plaque is the physical record of this propagation cycle. The size and clarity of the plaque can indicate the efficiency of the phage-host interaction sharp, clear plaques typically indicate highly specific, efficient lysis.
Grid squares with no plaque mean step 1 failed: the phage could not adsorb to the bacterial surface because the receptors didn’t match. Everything after that injection, replication, lysis never happened.
Myth vs. Fact: Common Misconceptions About Phage Typing
| Myth | Fact |
| Any phage can infect any bacterium | Bacteriophages are highly species-specific and often strain-specific determined by surface receptor matching |
| Plaques mean the phage “randomly attacked” the bacteria | Plaques are the result of highly specific molecular recognition adsorption is never random |
| No plaque means the phage is inactive or dead | No plaque means the phage is alive and functional the bacteria simply lack the matching surface receptors |
| Phage typing has been replaced by DNA methods | Phage typing is still used alongside molecular methods; in some settings it remains faster and more cost-effective |
| All bacterial strains of the same species respond to phages identically | Strains within the same species show distinct phage susceptibility profiles that the entire point of the method |
Real-World Applications: Why This Principle Actually Matters
The exercise isn’t just a classroom activity. The principle it demonstrates is applied in settings where lives depend on accurate bacterial identification.
Outbreak Investigations
When a foodborne illness outbreak occurs a Salmonella outbreak at a food processing plant, for example public health investigators need to determine whether cases in different locations are connected. Are patients sick from the same bacterial strain, or from different ones that happened to cause similar symptoms?
Phage typing answers that question. If bacterial isolates from multiple patients share the same phage type profile, they’re likely from a common source. Different profiles suggest unrelated infections. The 2008 Salmonella outbreak in the United States used exactly this kind of typing to trace infections across state lines.
Hospital Infection Control
Staphylococcus aureus is a major cause of hospital-acquired infections. When clusters of infections appear across different wards, infection control teams need to know: is this the same strain spreading through the hospital, or are patients acquiring different strains independently?
A landmark teaching exercise designed at a UK university uses simulated phage typing of S. aureus plates from three hypothetical hospital wards showing students how phage type profiles can identify whether an outbreak is common-source (same strain, same profile) or unrelated (different strains, different profiles).
Food Safety
Dairy, poultry, and meat processing industries use phage typing to monitor bacterial contamination and trace contamination events back to specific points in the production process. The same principle specific phage susceptibility patterns identifying specific bacterial strains enables traceability across complex supply chains.
See Also : Sovereign Foods Quality Control Job Matric Pass Fail Requirements
Phage Therapy Research
There is a growing interest in using bacteriophages therapeutically as targeted antimicrobial agents against antibiotic-resistant bacterial infections. Phage typing is foundational to this work because selecting the right phage for therapy requires knowing exactly which bacterial strain you’re targeting and which phage it’s susceptible to. As of 2026, research published in Nature Reviews Microbiology is actively exploring how machine learning models can predict phage-host specificity at the genomic level, building on the same biological principle the lab exercise demonstrates.
Comparison: Phage Typing vs. Modern Molecular Methods
| Feature | Phage Typing | Molecular Methods (PCR, WGS) |
| Speed | 24–48 hours | 4–72 hours depending on method |
| Cost | Low | Moderate to high |
| Equipment needed | Basic microbiology lab | Specialized molecular equipment |
| Discriminatory power | Good for major strains | Very high strain-level precision |
| Real-time applicability | High | Moderate |
| Complexity | Relatively simple | Technically demanding |
| Dependence on living phages | Yes | No |
| Best use case | Outbreak screening, field labs | Definitive strain characterization |
Both approaches are valid and often used together. Phage typing provides rapid preliminary classification; molecular methods provide definitive confirmation.
What Laboratory Experience Adds to Understanding
Having worked through phage typing exercises and studied their results, a consistent observation emerges that textbooks rarely capture: the most instructive moment isn’t seeing the plaques. It’s seeing the negative results.
Students expect the plaques clear zones are dramatic, easy to photograph, easy to point to. What takes longer to appreciate is the significance of the grid squares where nothing happens. An intact bacterial lawn under a phage suspension is not a failure. It’s a positive result, positive evidence of receptor mismatch. The exercise teaches that absence of lysis is data, not noise. That lesson generalizes beyond phage typing to the whole of experimental biology.
The second thing repeated practice reinforces: pattern interpretation requires consistency in technique. The same bacterial strain inoculated slightly too densely will produce faint plaques that could be misread as no lysis. The same phage applied at too low a concentration produces ambiguous results. Phage typing is a principle-driven technique, but it demands procedural precision. The principle only becomes visible when the technique is clean.
Conclusion
The phage typing exercise is one of microbiology’s most elegant teaching tools because it doesn’t just illustrate a principle it makes the principle physically observable.
When you describe how this exercise demonstrates the principle of phage typing, you’re describing a chain of events: a phage searches for a matching bacterial surface receptor, binds when it finds one, initiates the lytic cycle, and leaves behind a visible plaque as evidence. Where no plaque forms, the receptor didn’t match. The resulting pattern of presence and absence across the phage panel is a strain fingerprint, a biological signature written in lysis.
That principle bacteriophage host specificity connects a 100-year-old discovery in a Paris laboratory to 2026 research in machine learning-based phage-host prediction, to the infection control investigations happening right now in hospitals across the world.
Understanding it fully doesn’t just help you pass an exam. It gives you a real window into how microbiology actually works and why a virus smaller than most measurements your eye can detect has become one of science’s most precise tools for identifying the bacterial world.
What to do next: If you’re preparing for an exam, use the step-by-step breakdown and FAQ answers above as your revision framework. If you’re a lab instructor, the myth-vs-fact table and the section provide ready material for class discussion. And if you want to go deeper, the research on phage-host specificity prediction published in Nature Reviews Microbiology (2026) shows exactly where this foundational principle is headed next.
We hope this content helped you clearly understand the topic. visit Decretosupremo160 again for more practical, easy-to-follow guides. Don’t forget to share this with your friends so they can benefit too.
Frequently Asked Questions
The exercise shows that bacteriophages infect only bacteria with specific matching surface receptors. By applying a panel of known phages to a bacterial lawn and observing where plaques form, the exercise directly illustrates the host specificity that is the core principle of phage typing. The unique pattern of lysis identifies the bacterial strain.
plaque is a clear zone in a bacterial lawn where bacterial cells have been lysed by a bacteriophage. It forms when a phage successfully adsorbs to bacterial surface receptors, injects its genetic material, replicates inside the cell, and causes the cell to burst releasing new phages that infect and lyse neighboring bacteria, expanding the clear zone outward.
Because each bacteriophage only infects bacteria whose surface receptors match its tail fiber proteins, the lock-and-key specificity mechanism. Phages that don’t match the target bacterium’s receptors cannot adsorb and therefore cannot initiate infection. The bacterial lawn remains intact in those grid squares.
The pattern of positive results (plaques present) and negative results (no plaques) across the phage panel is compared to a reference chart of known phage type profiles. The specific combination of lysed and resistant results maps the bacterium to a documented phage type and its strain-level identity.
Yes, though its role has evolved. Phage typing is still used in public health laboratories for rapid bacterial strain identification during outbreak investigations. It is increasingly used alongside molecular methods like whole-genome sequencing, which provide greater discriminatory power. In 2026, research into phage-host specificity prediction using machine learning is actively expanding the scientific foundations the exercise demonstrates.
Teaching exercises most commonly use Staphylococcus aureus (relevant to hospital infection control) or Salmonella typhi (relevant to food safety and enteric disease). E. coli and Listeria are also used in research and food industry settings. The choice reflects both safety considerations for student labs and relevance to real-world epidemiological applications.